We’ve been working on building simulations in R over the past few articles. We are now to the point where we want to simulate data using regression models. To recap what we’ve done so far:
- Part 1 discussed the basic functions for simulating and sampling data in R.
- Part 2 walked us through how to perform bootstrap resampling and then simulate bivariate and multivariate distributions.
- Part 3 we worked making group comparisons by simulating thousands of t-tests
In an upcoming article we will use simulation to evaluate regression assumptions. But, before we do that, it might be useful to use a regression model reported in a research paper to simulate a data set. This can be useful if you are trying to better understand the paper (you can recalculate statistics and explore the data) or if you want to work on applying a different type of model approach to build hypotheses for future research.
As always, all code is freely available in the Github repository.
I’m going to take the regression model from a paper by Ferrari et al (2022), Performance and anthropometrics of classic powerlifters: Which characteristics matter? J Strength Cond Res., which looked to predict power lifting performance in male and female lifters.
The paper used several variables to try and predict Squat, Bench, Deadlift, and Powerlifting Total in raw power lifters. To keep this simple, I’ll focus on the model for predicting squat performance, which was:

The model had a standard error of estimate (SEE) of 20.3 and an r-squared of 0.83.
Getting the necessary info from the paper
Aside from the model coefficients, above, we also need to get the mean and standard deviation for each of the variables in the model (for both male and female lifters). We will use those parameters to simulate 1000 male and female lifters.
library(tidyverse)
# set the see for reproducibility
set.seed(82)
# sample size - simulate 1000 male and 1000 female lifters
n_males <- 1000
n_females <- 1000
# set the model coefficients for squats
intercept <- -145.7
years_exp_beta <- 4.3
pct_bf_beta <- -1.7
upper_arm_girth_beta <- 6
thigh_girth_beta <- 1.9
# Standard Error of Estimate (SEE) is sampled from a normal distribution with a mean = 0 and
# standard deviation of 20.3
see <- rnorm(n = n_males + n_females, mean = 0, sd = 20.3)
# get mean and sd for each of the model variables
years_exp_male <- rnorm(n = n_males, mean = 2.6, sd = 2.4)
years_exp_male <- ifelse(years_exp_male < 0, 0, years_exp_male)
years_exp_female <- rnorm(n = n_females, mean = 1.6, sd = 1.5)
years_exp_female <- ifelse(years_exp_female < 0, 0, years_exp_female)
pct_bf_male <- rnorm(n = n_males, mean = 11.1, sd = 3.8)
pct_bf_female <- rnorm(n = n_females, mean = 21.7, sd = 5.4)
upper_arm_girth_male <- rnorm(n = n_males, mean = 35.6, sd = 2.8)
upper_arm_girth_female <- rnorm(n = n_females, mean = 29.5, sd = 3.1)
thigh_girth_male <- rnorm(n = n_males, mean = 61.1, sd = 5.5)
thigh_girth_female <- rnorm(n = n_females, mean = 56.1, sd = 4.9)
# put the simulated data into a data frame
dat <- data.frame( gender = c(rep("male", times = n_males), rep("female", times = n_females)), years_exp = c(years_exp_male, years_exp_female), pct_bf = c(pct_bf_male, pct_bf_female), upper_arm_girth = c(upper_arm_girth_male, upper_arm_girth_female), thigh_girth = c(thigh_girth_male, thigh_girth_female) ) dat %>%
head()
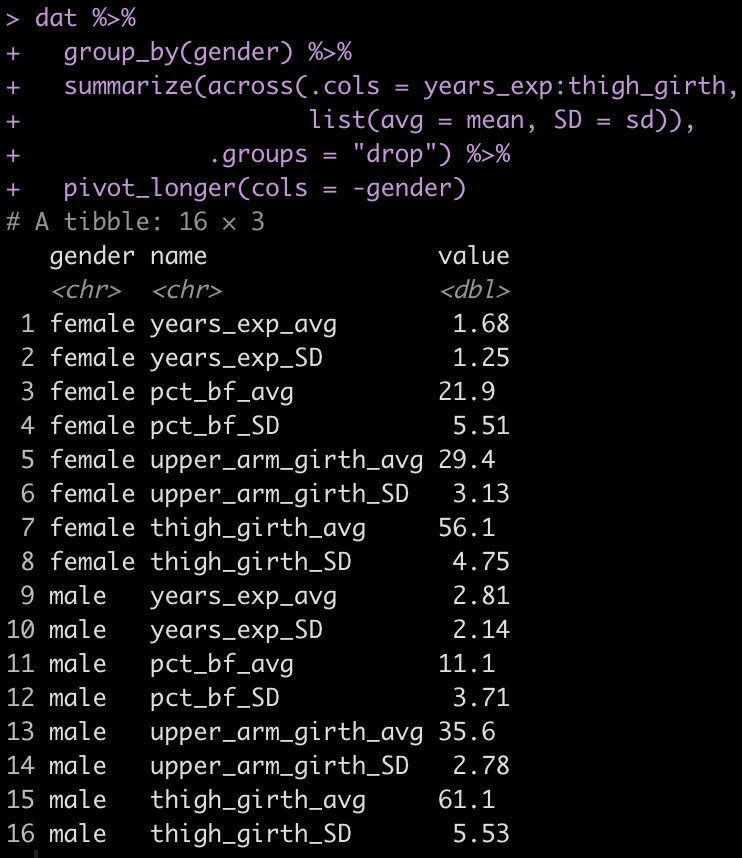
As a sanity check, we can quickly check the mean and standard deviation of each variable to ensure the simulation appears as we intended it to.
## check means and standard deviations of the simulation
dat %>%
group_by(gender) %>%
summarize(across(.cols = years_exp:thigh_girth,
list(avg = mean, SD = sd)),
.groups = "drop") %>%
pivot_longer(cols = -gender)
Estimate squat performance using the model
Next, we use the values that we simulated and the model coefficients to simulate the outcome of interest (Squat performance).
# estimate squat performance
dat$squat <- with(dat, intercept + years_exp_beta*years_exp + pct_bf_beta*pct_bf + upper_arm_girth_beta*upper_arm_girth + thigh_girth_beta*thigh_girth + see) dat %>%
head()
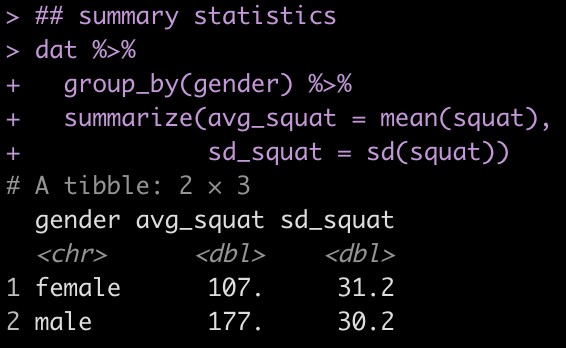
## summary statistics
dat %>%
group_by(gender) %>%
summarize(avg_squat = mean(squat),
sd_squat = sd(squat))
# plots
hist(dat$squat)
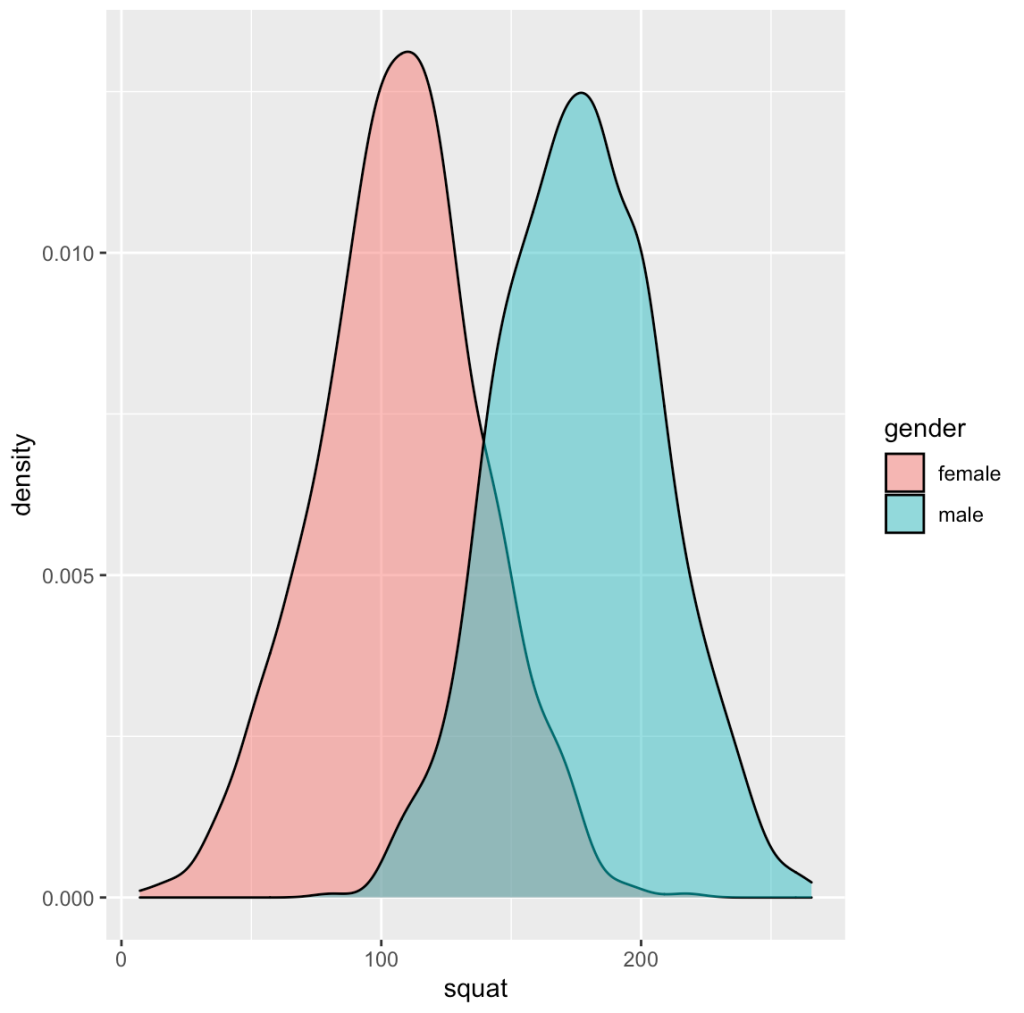
dat %>%
ggplot(aes(x = squat, fill = gender)) +
geom_density(alpha = 0.5)
Look at the regression model
Now we fit a regression model with our simulated data.
fit_lm <- lm(squat ~ years_exp + pct_bf + upper_arm_girth + thigh_girth, data = dat)
summary(fit_lm)
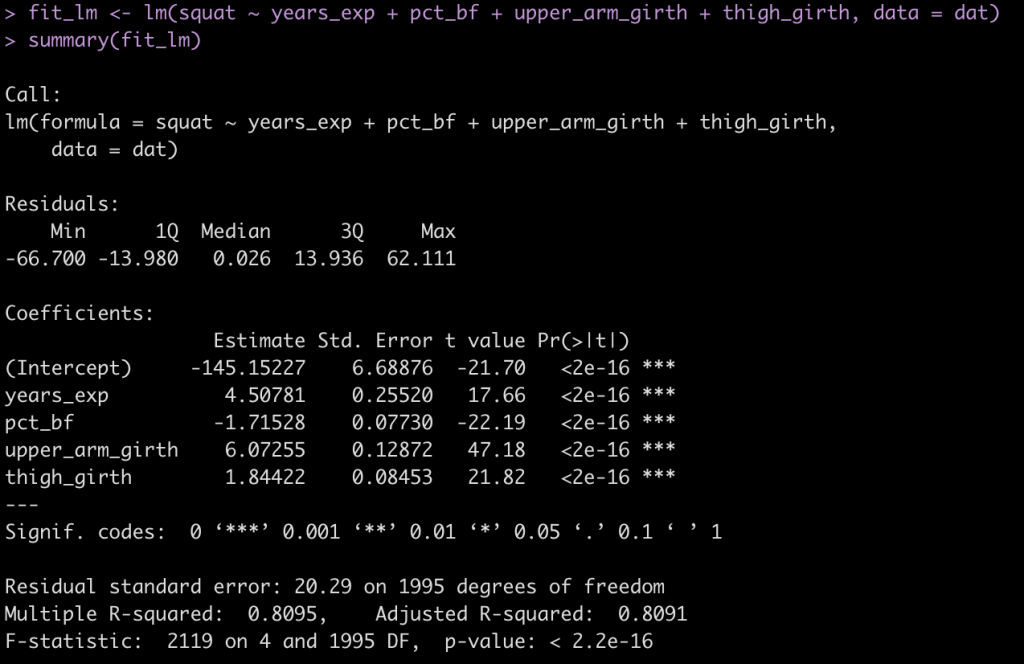
- The coefficients are similar to those in the paper (which makes sense because we coded them this way.
- The standard error (Residual standard error) is 20.3, as expected.
- The r-squared for the model is 0.81, close to what was observed in the paper.
Making a better simulation
One thing to notice is the mean and standard deviation of the squat in our simulation is a bit less compared to what is reported for the population in the paper.

Our estimates are a little low relative to what was reported in the paper. The model above still works because we constructed it with the regression coefficients and variable parameters in the model, so we can still play with the data and learn something. But, the estimated squats might be a little low because the anthropometric variables in the model (BF%, Upper Arm Girth, and Thigh Girth) are in some way going to be correlated with each other. So, we could make this simulation more informative by simulating those variables from a multivariate normal distribution, as we did in Part 3.
To start, we load the mvtnorm package and set up a vector of mean values for each variable. We will construct a vector for males and females separately. We will use the mean values for each variable reported in the paper.
library(mvtnorm)
## Order of means is: BF%, Upper Arm Girth, Thigh Girth
male_means <- c(11.1, 35.6, 61.1)
female_means <- c(21.7, 29.5, 56.1)
Next, we need to construct a correlation matrix between these three variables. Again, we will create a correlation matrix for males and females, separately. I’m not sure of the exact correlation between these variables, so I’ll just estimate what I believe it to be. For example, the correlations probably aren’t 0.99 but they also probably aren’t below 0.6. I’m also unsure how these correlations might differ between the two genders. Thus, I’ll keep the same correlation matrix for both. To keep it simple, I’ll set the correlation between BF% and upper arm and thigh girth at 0.85 and the correlation between upper arm girth and thigh girth to be 0.9. In theory, we could consult some scientific literature on these things and attempt to construct more plausible correlations.
## Create correlation matrices
# males
male_r_matrix <- matrix(c(1, 0.85, 0.85,
0.85, 1, 0.9,
0.85, 0.9, 1),
nrow = 3, ncol = 3,
dimnames = list(c("bf_pct", "upper_arm_girth", "thigh_girth"),
c("bf_pct", "upper_arm_girth", "thigh_girth")))
male_r_matrix
# females
female_r_matrix <- matrix(c(1, 0.85, 0.85,
0.85, 1, 0.9,
0.85, 0.9, 1),
nrow = 3, ncol = 3,
dimnames = list(c("bf_pct", "upper_arm_girth", "thigh_girth"),
c("bf_pct", "upper_arm_girth", "thigh_girth")))
female_r_matrix
Now we will create 1000 simulations from a multivariate normal distribution for both males and females and then row bind them together into a single big data frame.
## simulate 1000 new x, y, and z variables using the mvtnorm package
set.seed(777)
male_sim <- rmvnorm(n = n_males, mean = male_means, sigma = male_r_matrix) %>%
as.data.frame() %>%
setNames(c("pct_bf", "upper_arm_girth", "thigh_girth"))
female_sim <- rmvnorm(n = n_females, mean = female_means, sigma = female_r_matrix) %>%
as.data.frame() %>%
setNames(c("pct_bf", "upper_arm_girth", "thigh_girth"))
head(male_sim)
head(female_sim)
## put the two simulated data frames together
multi_sims <- bind_rows(male_sim, female_sim) multi_sims %>%
head()
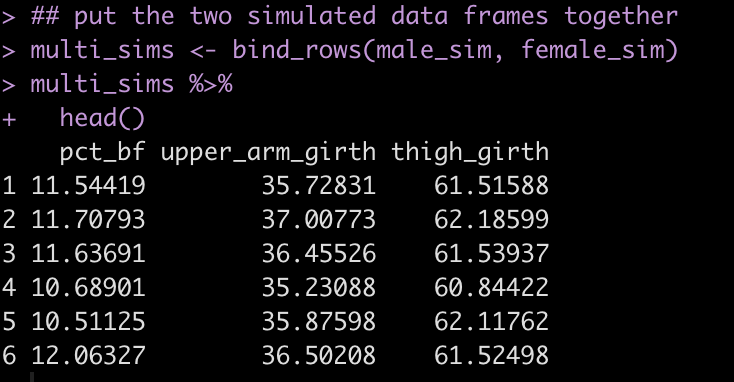
Finally, one last thing we’ll change is our simulation of years of experience. This variable is not a normally distributed variable because it is truncated at 0. Above, in our first simulation, we attempted to solve this with the ifelse() expression to assign any simulated value less than 0 to 0. Here, I’ll just get the quantiles of the simulated years of experience so that I have an idea of a plausible upper end of experience for the population used in this paper. Then, instead of simulating from a normal distribution I’ll do a random draw from a uniform distribution from 0 to the respective upper end for the male and female groups.
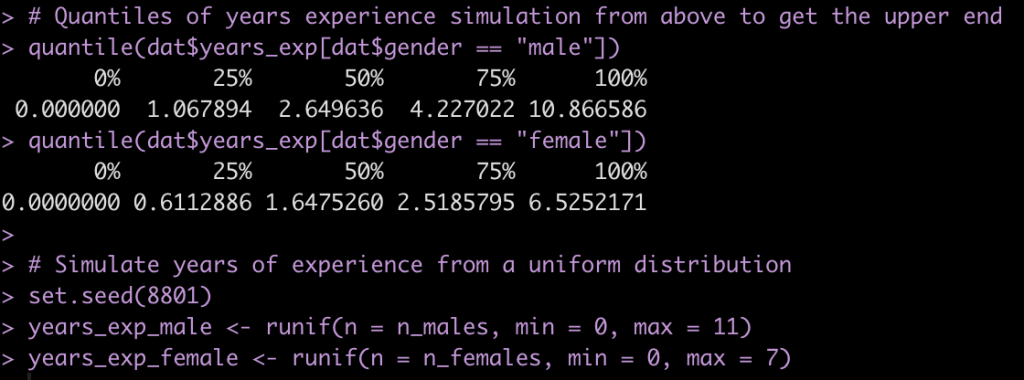
Now take the newly simulated variables and create a new data frame.
## new data frame
new_dat <- data.frame( gender = c(rep("male", times = n_males), rep("female", times = n_females)), years_exp = c(years_exp_male, years_exp_female) ) %>%
bind_cols(multi_sims)
new_dat %>%
head()
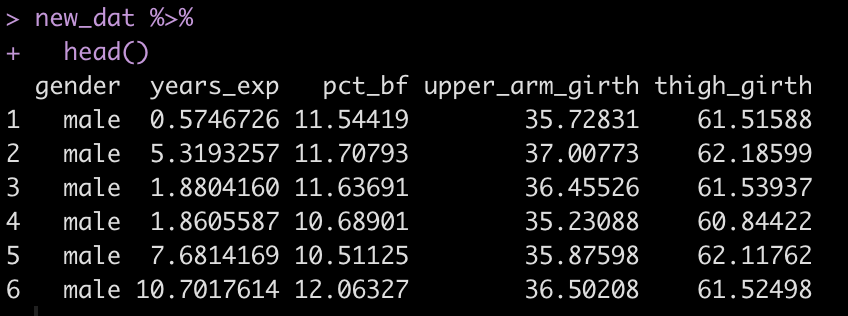
Finally, go through the steps we did above to estimate the squat from our four variables and the beta coefficients from the paper.
# estimate squat performance
new_dat$squat <- with(new_dat, intercept + years_exp_beta*years_exp + pct_bf_beta*pct_bf + upper_arm_girth_beta*upper_arm_girth + thigh_girth_beta*thigh_girth + see) new_dat %>%
head()
## summary statistics
new_dat %>%
group_by(gender) %>%
summarize(avg_squat = mean(squat),
sd_squat = sd(squat))
# plots
hist(new_dat$squat)
new_dat %>%
ggplot(aes(x = squat, fill = gender)) +
geom_density(alpha = 0.5)
Look at the new regression model
fit_lm_new <- lm(squat ~ years_exp + pct_bf + upper_arm_girth + thigh_girth, data = new_dat)
summary(fit_lm_new)
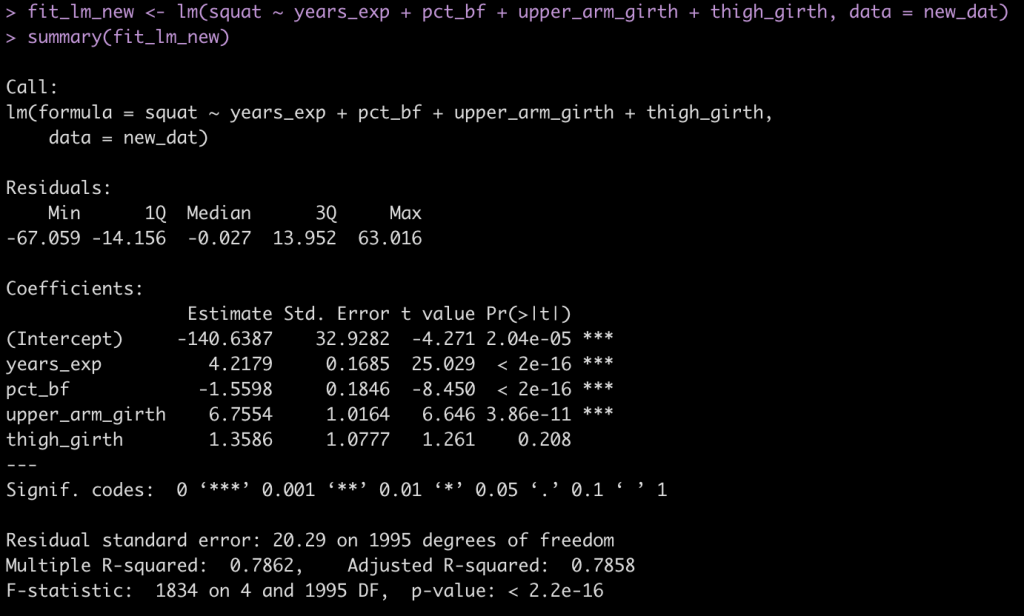
Compare the squat simulated data to that which was reported in the paper
new_dat %>%
group_by(gender) %>%
summarize(avg_squat_sim = mean(squat),
sd_squat_sim = sd(squat)) %>%
mutate(avg_squat_paper = c(118.3, 196.1),
sd_squat_paper = c(26.6, 37.9))

Now we have mean values much closer to those observed in the population of the study. Our standard deviations differ from the study because, if you recall, the standard error of the estimate was 20.3 for the regression model. The model in the study did not include gender as an independent variable. This to me is a little strange and in the discussion of the paper the authors’ also indicate that it was strange to them too. However, the approach the authors’ took to building this deemed gender unnecessary as a predictor variable. Consequently, our simulation has a similar standard deviation in estimate squat for both the male and female populations. However, we now have a data set that is relatively close to what was observed in the paper and can therefore proceed with conducting other statistical tests or building other models.
Wrapping Up
We’ve come a long way in our simulation journey, from randomly drawing single vectors of data to now building up entire data sets using regression models and simulating data from research studies. Next time we will use simulation to explore regression assumptions.
As always, all code is freely available in the Github repository.