I’ve talked a lot in this blog about making predictions (see HERE, HERE, and HERE) as well as the difference between confidence intervals and prediction intervals and why you’d use one over the other (see HERE). Tonight I was having a discussion with a colleague about some models he was working on and he was building some confidence intervals around his predictions. That got me to thinking about the various ways we can code confidence intervals, quantile intervals, and prediction intervals in R. So, I decided to put together this quick tutorial to provide a few different ways of constructing these values (after all, unless we can calculate the uncertainty in our predictions, point estimate predictions are largely useless on their own).
The full code is available on my GITHUB page.
Load packages, get data, and fit regression model
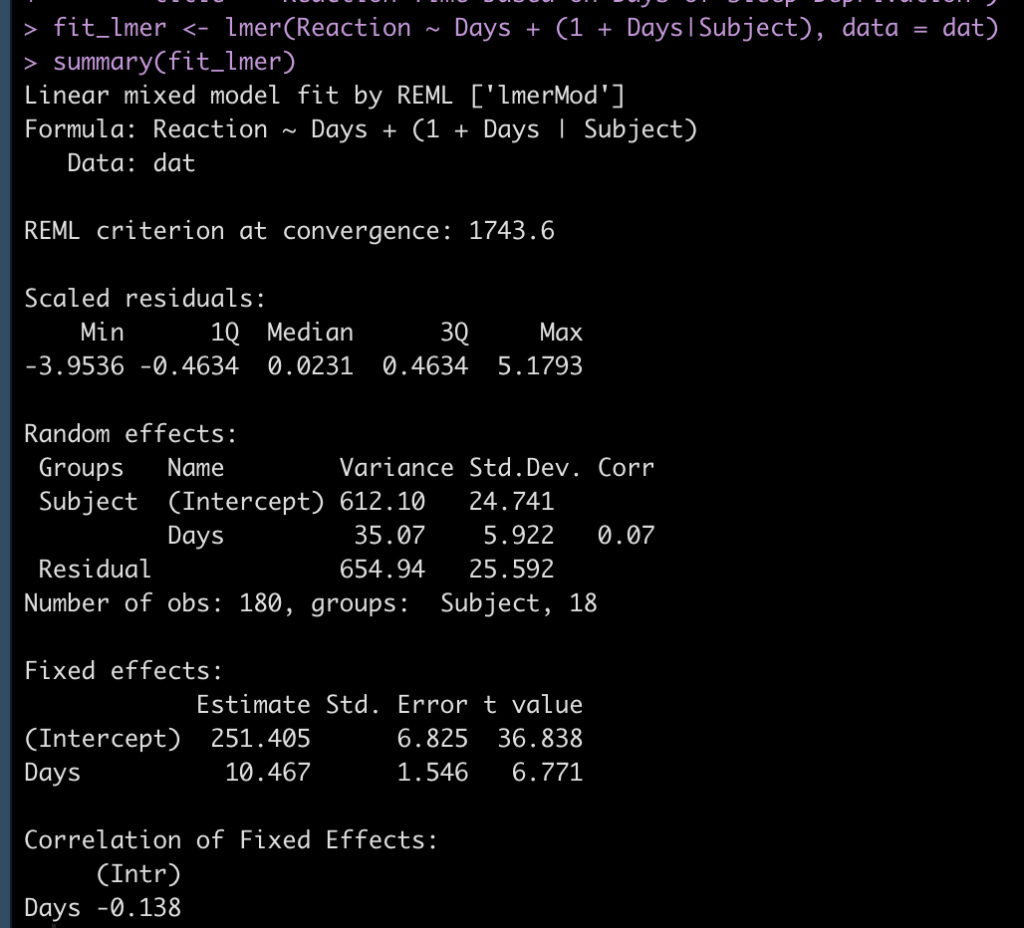
The only package we will need is {tidyverse}, the data will be the mtcars dataset and the model will be a linear regression which attempts to predict mpg from wr and carb.
## Load packages library(tidyverse) theme_set(theme_classic()) ## Get data d <- mtcars d %>% head() ## fit model fit_lm <- lm(mpg ~ wt + carb, data = d) summary(fit_lm)
Get some data to make predictions on
We will just grab a random sample of 5 rows from the original data set and use that to make some predictions on.
## Get a few rows to make predictions on set.seed(1234) d_sample <- d %>% sample_n(size = 5) %>% select(mpg, wt, carb) d_sample
Confidence Intervals with the predict() function
Using preidct() we calculate the predicted value with 95% Confidence Intervals.
## 95% Confidence Intervals
d_sample %>%
bind_cols(
predict(fit_lm, newdata = d_sample, interval = "confidence", level = 0.95)
)

Calculate confidence intervals by hand
Instead of using the R function, we can calculate the confidence intervals by hand (and obtain the same result).
## Calculate the 95% confidence interval by hand
level <- 0.95
alpha <- 1 - (1 - level) / 2
t_crit <- qt(p = alpha, df = fit_lm$df.residual)
d_sample %>%
mutate(pred = predict(fit_lm, newdata = .),
se_pred = predict(fit_lm, newdata = ., se = TRUE)$se.fit,
cl95 = t_crit * se_pred,
lwr = pred - cl95,
upr = pred + cl95)

Calculate confidence intervals with the qnorm() function
Above, we calculated a 95% t-critical value for the degrees of freedom of our model. Alternatively, we could calculate 95% confidence intervals using the standard z-critical value for 95%, 1.96, which we obtain with the qnorm() function.
d_sample %>%
mutate(pred = predict(fit_lm, newdata = .),
se_pred = predict(fit_lm, newdata = ., se = TRUE)$se.fit,
lwr = pred + qnorm(p = 0.025, mean = 0, sd = 1) * se_pred,
upr = pred + qnorm(p = 0.975, mean = 0, sd = 1) * se_pred)
Calculate quantile intervals via simulation
Finally, we can calculate quantile intervals by simulating predictions using the predicted value and standard error for each of the observations. We simulate 1000 times from a normal distribution and then use the quantile() function to get our quantile intervals.
If all we care about is a predicted value and the lower and upper intervals, we can use the rowwise() function to indicate that we are going to do a simulation for each row and then store the end result (our lower and upper quantile intervals) in a new column.
## 95% Quantile Intervals via Simulation
d_sample %>%
mutate(pred = predict(fit_lm, newdata = .),
se_pred = predict(fit_lm, newdata = ., se = TRUE)$se.fit) %>%
rowwise() %>%
mutate(lwr = quantile(rnorm(n = 1000, mean = pred, sd = se_pred), probs = 0.025),
upr = quantile(rnorm(n = 1000, mean = pred, sd = se_pred), probs = 0.975))
While that is useful, there might be times where we want to extract the full simulated distribution. We can create a simulated distribution (1000 simulations) for each of the 5 observations using a for() loop.
## 95% quantile intervals via Simulation with full distribution
N <- 1000
pred_sim <- list()
set.seed(8945)
for(i in 1:nrow(d_sample)){
pred <- predict(fit_lm, newdata = d_sample[i, ])
se_pred <- predict(fit_lm, newdata = d_sample[i, ], se = TRUE)$se.fit
pred_sim[[i]] <- rnorm(n = N, mean = pred, sd = se_pred)
}
sim_df <- tibble( sample_row = rep(1:5, each = N), pred_sim = unlist(pred_sim) )
sim_df %>%
head()

Next we summarize the simulation for each observation.
# get predictions and quantile intervals
sim_df %>%
group_by(sample_row) %>%
summarize(pred = mean(pred_sim),
lwr = quantile(pred_sim, probs = 0.025),
upr = quantile(pred_sim, probs = 0.975)) %>%
mutate(sample_row = rownames(d_sample))

We can then plot the entire posterior distribution for each observation.
# plot the predicted distributions
sim_df %>%
mutate(actual_value = rep(d_sample$mpg, each = N),
sample_row = case_when(sample_row == 1 ~ "Hornet 4 Drive",
sample_row == 2 ~ "Toyota Corolla",
sample_row == 3 ~ "Honda Civic",
sample_row == 4 ~ "Ferrari Dino",
sample_row == 5 ~ "Pontiac Firebird")) %>%
ggplot(aes(x = pred_sim)) +
geom_histogram(color = "white",
fill = "light grey") +
geom_vline(aes(xintercept = actual_value),
color = "red",
size = 1.2,
linetype = "dashed") +
facet_wrap(~sample_row, scale = "free_x") +
labs(x = "Predicted Simulation",
y = "count",
title = "Predicted Simulation with actual observation (red line)",
subtitle = "Note that the x-axis are specific to that simulation and not the same")
Prediction Intervals with the predict() function
Next we turn attention to prediction intervals, which will be wider than the confidence intervals because they are incorporating additional uncertainty.
The predict() function makes calculating prediction intervals very convenient.
## 95% Prediction Intervals
d_sample %>%
bind_cols(
predict(fit_lm, newdata = d_sample, interval = "predict", level = 0.95)
)
Prediction Intervals from a simulated distribution
Similar to how we simulated a distribution for calculating quantile intervals, above, we will perform the same procedure here. The difference is that we need to get the residual standard error (RSE) from our model as we need to add this additional piece of uncertainty (on top of the predicted standard error) to each of the simulated predictions.
## 95% prediction intervals from a simulated distribution
# store the model residual standard error
sigma <- summary(fit_lm)$sigma
# run simulation
N <- 1000
pred_sim2 <- list()
set.seed(85)
for(i in 1:nrow(d_sample)){
pred <- predict(fit_lm, newdata = d_sample[i, ])
se_pred <- predict(fit_lm, newdata = d_sample[i, ], se = TRUE)$se.fit
pred_sim2[[i]] <- rnorm(n = N, mean = pred, sd = se_pred) + rnorm(n = N, mean = 0, sd = sigma)
}
# put results in a data frame
sim_df2 <- tibble( sample_row = rep(1:5, each = N), pred_sim2 = unlist(pred_sim2) )
sim_df2 %>%
head()
We summarize our predictions and their intervals.
# get predictions and intervals
sim_df2 %>%
group_by(sample_row) %>%
summarize(pred = mean(pred_sim2),
lwr = quantile(pred_sim2, probs = 0.025),
upr = quantile(pred_sim2, probs = 0.975)) %>%
mutate(sample_row = rownames(d_sample))
Finally, we plot the simulated distributions for each of the observations.
Wrapping Up
Uncertainty is important to be aware of and convey whenever you share your predictions. The point estimate prediction is one a single value of many plausible values given the data generating process. This article provided a few different approaches for calculating uncertainty intervals. The full code is available on my GITHUB page.